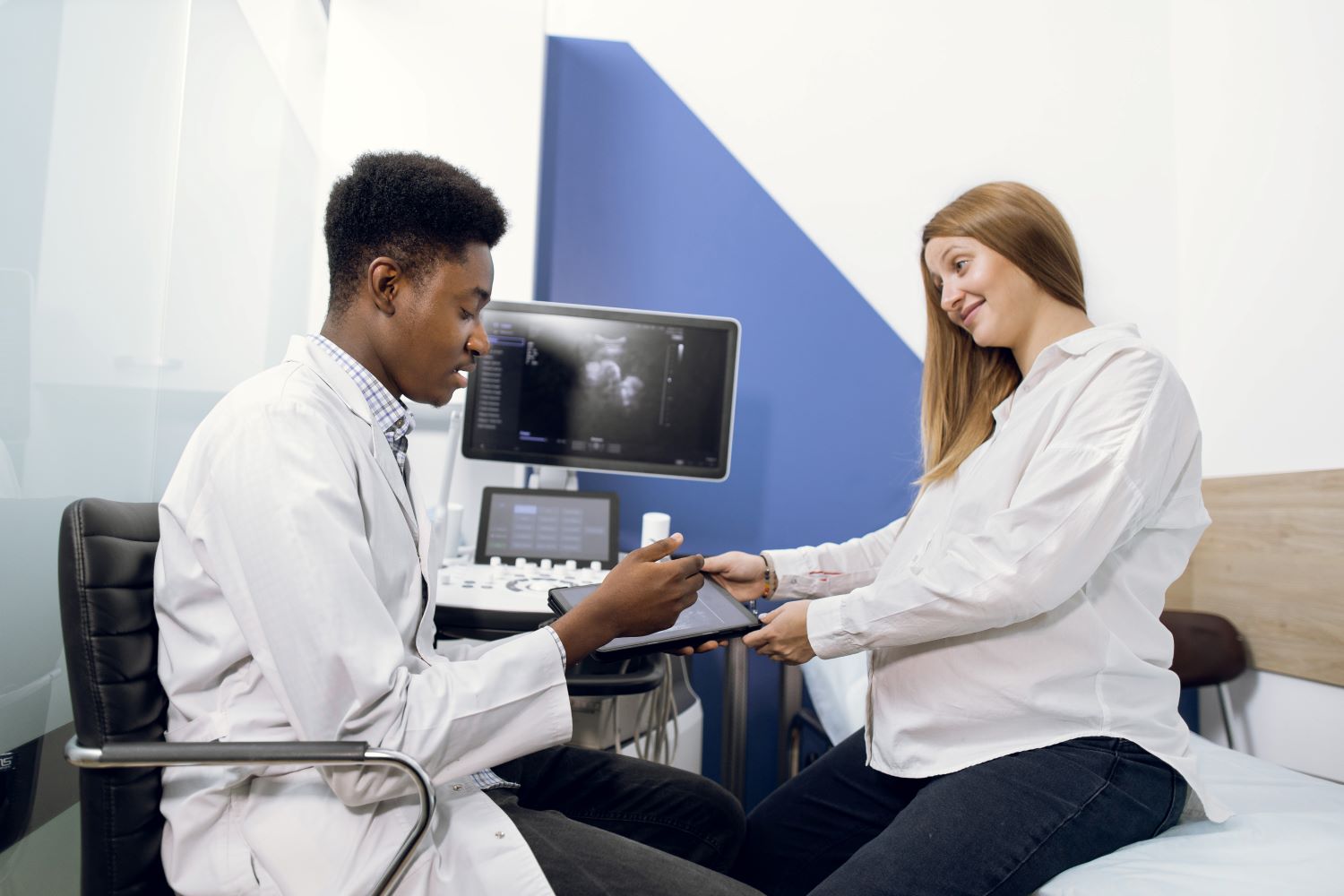
หนึ่งในคำถามที่พบบ่อยที่สุดสำหรับผู้ที่ต้องการเรียนภาษาอังกฤษออนไลน์ตัวต่อตัวคือ จะเลือกเรียนพิเศษภาษาอังกฤษแบบตัวต่อตัวแบบตัวต่อตัวหรือแบบกลุ่ม ทั้งสองรูปแบบมีข้อดีและข้อเสีย และการเลือกระหว่างรูปแบบทั้งสองจะขึ้นอยู่กับความต้องการและความชอบของแต่ละคน ในบทความนี้ เราจะประเมินความแตกต่างระหว่างการสอนและการเรียนภาษาอังกฤษออนไลน์ตัวต่อตัวกับการสอนแบบกลุ่มเพื่อพิจารณาว่าแบบใดเหมาะสมกับการเรียนภาษาอังกฤษมากกว่ากัน ข้อดีของการสอนภาษาอังกฤษออนไลน์แบบตัวต่อตัว: ความใส่ใจส่วนบุคคล: ในการเรียนภาษาอังกฤษออนไลน์ตัวต่อตัว นักเรียนจะได้รับความสนใจเป็นรายบุคคลเนื่องจากผู้สอนเน้นเฉพาะความต้องการในการเรียนรู้ของพวกเขาเท่านั้น ผู้สอนสามารถปรับแต่งหลักสูตรให้เหมาะกับระดับความสามารถและวัตถุประสงค์ของนักเรียน และพัฒนาหลักสูตรเฉพาะสำหรับพวกเขา แผนการสอนที่ปรับให้เหมาะกับคุณ: รูปแบบการสอนการเรียนภาษาอังกฤษออนไลน์ตัวต่อตัวเป็นแนวทางที่มุ่งเน้นและปรับให้เหมาะกับความต้องการด้านภาษาของนักเรียนมากขึ้น ผู้สอนสามารถพัฒนาแผนการสอนที่รวมถึงความสนใจและเป้าหมายของนักเรียน ความยืดหยุ่นที่เพิ่มขึ้น: การสอนออนไลน์จะสามารถมอบความยืดหยุ่นในการจัดตารางการประชุมมากกว่าการเรียนรู้ในห้องเรียนแบบดั้งเดิม และดำเนินการได้ในเวลาและสถานที่ที่ต้องการของนักเรียน ข้อเสียเรียนภาษาอังกฤษออนไลน์ตัวต่อตัว แพง: […]
Read more →